User Guide
Pan-Cancer Prediction Model
Universal prediction model applicable to multiple cancer types, supporting single sequence prediction and batch CSV file prediction.
Single Sequence Prediction
Input Format:
seq_a:MSGRGKQGGKARAKAKTRSS...,seq_b:MSGRGKQGGKARAKAKTRSS...- • seq_a: Wild-type protein sequence
- • seq_b: Mutant protein sequence
- • Both sequences must be the same length with only a single variant position
In the Pan-Cancer Prediction Model section, click Single Sequence Prediction, enter the wild-type and mutant protein sequences in the input box, then click Submit and wait for the results to appear.

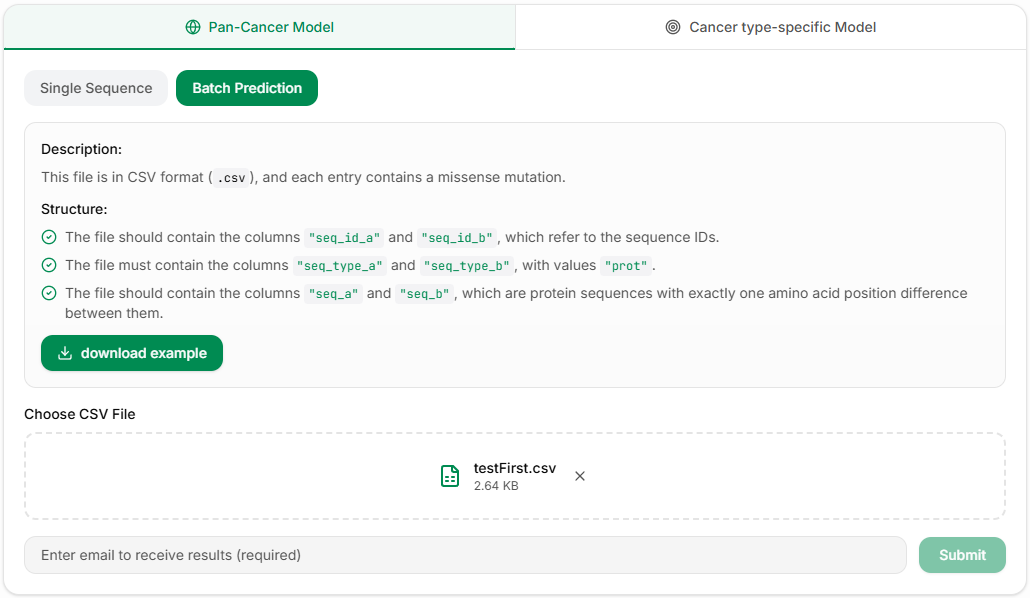
Batch Prediction (CSV File Upload)
For large amounts of data, you can use CSV file batch upload. File format requirements:
| Column Name | Description |
|---|---|
| seq_id_a, seq_id_b | Unique sequence identifiers |
| seq_type_a, seq_type_b | Sequence type, value is "prot" |
| seq_a, seq_b | Protein sequences, both must be the same length with only a single variant position |
Note: Batch prediction requires a valid email address, results will be sent via email.
In the Pan-Cancer Prediction Model section, click Batch Prediction, upload a CSV file that strictly follows the above requirements, and enter your email to receive the processed results.
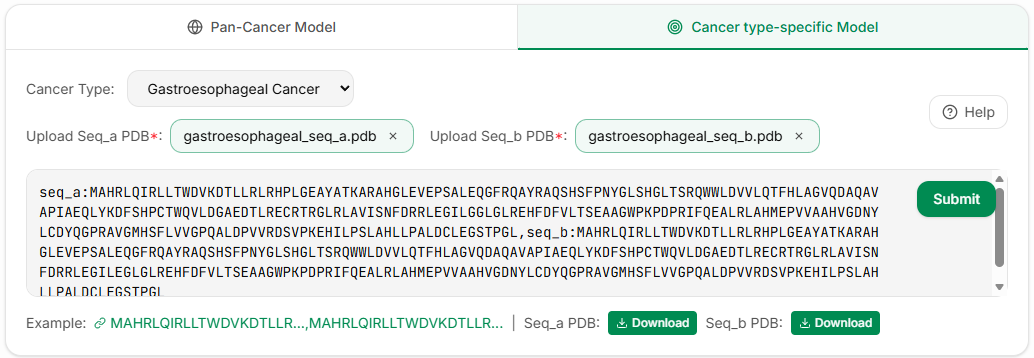
Cancer type-specific Prediction Model
For specific cancer types, providing more accurate cancer-specific predictions by integrating protein sequence semantics and spatial structural features. Currently supports the following cancer types:
Gastroesophageal Cancer
Leukemia
Lung Cancer
Usage Instructions
- • Select the target cancer type from the dropdown list
- • Input format follows seq_a (wild-type sequence) and seq_b (mutant sequence) corresponding format
- • The model only supports single amino acid substitution prediction, please ensure only a single residue difference between the two sequences
- • PDB format files must be uploaded, the system will extract protein structural features and combine them with sequence features for enhanced prediction
In the Cancer type-specific Prediction Model section, click the dropdown menu to select a specific cancer type, enter the wild-type and mutant protein sequences and upload the corresponding PDB files, then click Submit and wait for the results to appear.
Prediction Results Explanation
- • Single Prediction (Pan-Cancer/Cancer type-specific):After submitting a task, results will be displayed directly on the page
- • Batch Prediction (Pan-Cancer Model only):After submitting a task, results will be sent to your provided email
- • Processing time depends on task complexity, please wait patiently
- • The predicted score range is from 0 to 1. The higher the score, the greater the possibility that the mutation is a driver mutation.
- • The system's default threshold for determination is 0.5. That is, when the score is ≥ 0.5, it is determined as a driver mutation; when the score is < 0.5, it is determined as a passenger mutation
Driver mutations are mutations that promote tumor development and progression.
Passenger mutations are neutral mutations that have no significant role in promoting tumor development.
Important Notes
- • Please ensure the input sequence format is correct, otherwise prediction may fail
- • Both sequences must be the same length with only a single amino acid position difference
- • Batch prediction requires a valid email address to receive results
- • If you encounter any issues, please contact zhenyuyue@ahau.edu.cn
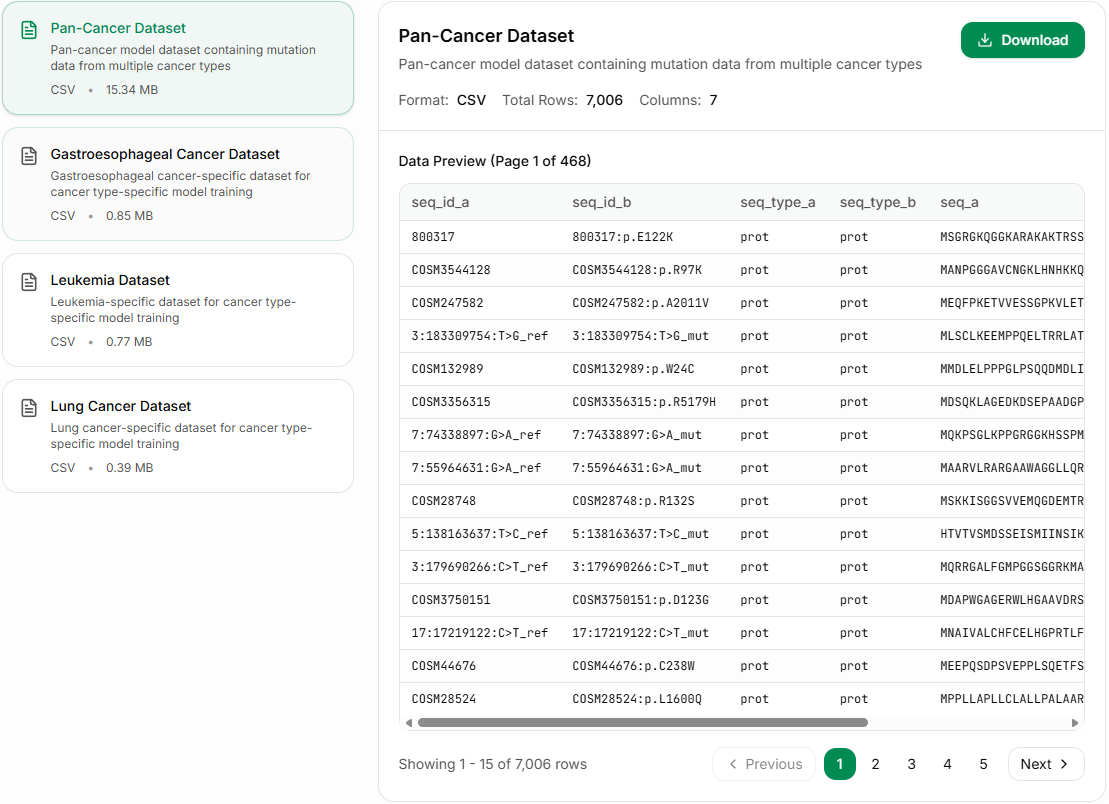
Datasets
These are the datasets used by this model. You can view or download them on this website.
Contact Us
If you encounter any issues during use, or have suggestions for improvement, please feel free to contact us at any time.